CNN
—
In February 2016, infectious disease epidemiologist Steffanie Strathdee was holding her dying husband’s hand, watching him lose an exhausting fight against a deadly superbug infection.
After months of ups and downs, doctors had just told her that her husband, Tom Patterson, was too racked with bacteria to live.
“I told him, ‘Honey, we’re running out of time. I need to know if you want to live. I don’t even know if you can hear me, but if you can hear me and you want to live, please squeeze my hand.’
“All of a sudden, he squeezed really hard. And I thought, ‘Oh, great!’ And then I’m thinking, ‘Oh, crap! What am I going to do?’”
What she accomplished next could easily be called miraculous. First, Strathdee found an obscure treatment that offered a glimmer of hope — fighting superbugs with phages, viruses created by nature to eat bacteria.
Then she convinced phage scientists around the country to hunt and peck through molecular haystacks of sewage, bogs, ponds, the bilge of boats and other prime breeding grounds for bacteria and their viral opponents. The impossible goal: quickly find the few, exquisitely unique phages capable of fighting a specific strain of antibiotic-resistant bacteria literally eating her husband alive.
Next, the US Food and Drug Administration had to greenlight this unproven cocktail of hope, and scientists had to purify the mixture so that it wouldn’t be deadly.
Yet just three weeks later, Strathdee watched doctors intravenously inject the mixture into her husband’s body — and save his life.
Their story is one of unrelenting perseverance and unbelievable good fortune. It’s a glowing tribute to the immense kindness of strangers. And it’s a story that just might save countless lives from the growing threat of antibiotic-resistant superbugs — maybe even your own.
“It’s estimated that by 2050, 10 million people per year — that’s one person every three seconds — is going to be dying from a superbug infection,” Strathdee told an audience at Life Itself, a 2022 health and wellness event presented in partnership with CNN.
“I’m here to tell you that the enemy of my enemy can be my friend. Viruses can be medicine.”
How this ‘perfect predator’ saved his life after nine months in the hospital
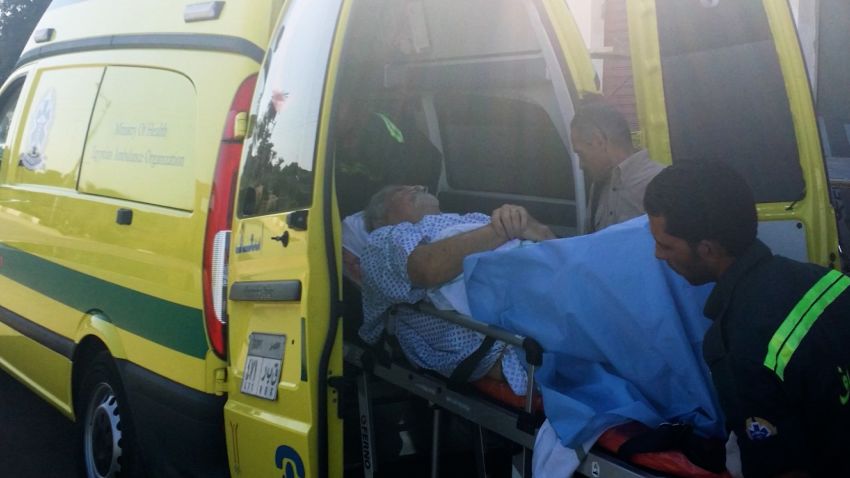
During a Thanksgiving cruise on the Nile in 2015, Patterson was suddenly felled by severe stomach cramps. When a clinic in Egypt failed to help his worsening symptoms, Patterson was flown to Germany, where doctors discovered a grapefruit-size abdominal abscess filled with Acinetobacter baumannii, a virulent bacterium resistant to nearly all antibiotics.
Found in the sands of the Middle East, the bacteria were blown into the wounds of American troops hit by roadside bombs during the Iraq War, earning the pathogen the nickname “Iraqibacter.”
“Veterans would get shrapnel in their legs and bodies from IED explosions and were medevaced home to convalesce,” Strathdee told CNN, referring to improvised explosive devices. “Unfortunately, they brought their superbug with them. Sadly, many of them survived the bomb blasts but died from this deadly bacterium.”
Today, Acinetobacter baumannii tops the World Health Organization’s list of dangerous pathogens for which new antibiotics are critically needed.
“It’s something of a bacterial kleptomaniac. It’s really good at stealing antimicrobial resistance genes from other bacteria,” Strathdee said. “I started to realize that my husband was a lot sicker than I thought and that modern medicine had run out of antibiotics to treat him.”
With the bacteria growing unchecked inside him, Patterson was soon medevaced to the couple’s hometown of San Diego, where he was a professor of psychiatry and Strathdee was the associate dean of global health sciences at the University of California, San Diego.
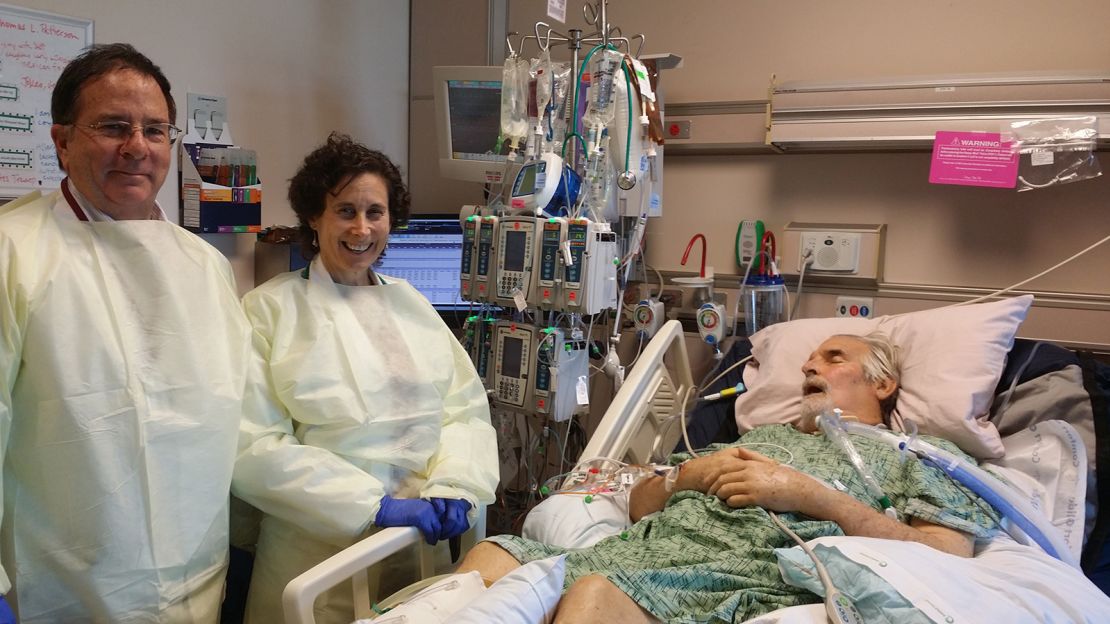
“Tom was on a roller coaster — he’d get better for a few days, and then there would be a deterioration, and he would be very ill,” said Dr. Robert “Chip” Schooley, a leading infectious disease specialist at UC San Diego who was a longtime friend and colleague. As weeks turned into months, “Tom began developing multi-organ failure. He was sick enough that we could lose him any day.”
After that reassuring hand squeeze from her husband, Strathdee sprang into action. Scouring the internet, she had already stumbled across a study by a Tbilisi, Georgia, researcher on the use of phages for treatment of drug-resistant bacteria.
A phone call later, Strathdee discovered phage treatment was well established in former Soviet bloc countries but had been discounted long ago as “fringe science” in the West.
“Phages are everywhere. There’s 10 million trillion trillion — that’s 10 to the power of 31 — phages that are thought to be on the planet,” Strathdee said. “They’re in soil, they’re in water, in our oceans and in our bodies, where they are the gatekeepers that keep our bacterial numbers in check. But you have to find the right phage to kill the bacterium that is causing the trouble.”
Buoyed by her newfound knowledge, Strathdee began reaching out to scientists who worked with phages: “I wrote cold emails to total strangers, begging them for help,” she said at Life Itself.
One stranger who quickly answered was Texas A&M University biochemist Ryland Young. He’d been working with phages for over 45 years.
“You know the word persuasive? There’s nobody as persuasive as Steffanie,” said Young, a professor of biochemistry and biophysics who runs the lab at the university’s Center for Phage Technology. “We just dropped everything. No exaggeration, people were literally working 24/7, screening 100 different environmental samples to find just a couple of new phages.”
While the Texas lab burned the midnight oil, Schooley tried to obtain FDA approval for the injection of the phage cocktail into Patterson. Because phage therapy has not undergone clinical trials in the United States, each case of “compassionate use” required a good deal of documentation. It’s a process that can consume precious time.
But the woman who answered the phone at the FDA said, “‘No problem. This is what you need, and we can arrange that,’” Schooley recalled. “And then she tells me she has friends in the Navy that might be able to find some phages for us as well.”
In fact, the US Naval Medical Research Center had banks of phages gathered from seaports around the world. Scientists there began to hunt for a match, “and it wasn’t long before they found a few phages that appeared to be active against the bacterium,” Strathdee said.
Back in Texas, Young and his team had also gotten lucky. They found four promising phages that ravaged Patterson’s antibiotic-resistant bacteria in a test tube. Now the hard part began — figuring out how to separate the victorious phages from the soup of bacterial toxins left behind.
“You put one virus particle into a culture, you go home for lunch, and if you’re lucky, you come back to a big shaking, liquid mess of dead bacteria parts among billions and billions of the virus,” Young said. “You want to inject those virus particles into the human bloodstream, but you’re starting with bacterial goo that’s just horrible. You would not want that injected into your body.”
Purifying phage to be given intravenously was a process that no one had yet perfected in the US, Schooley said, “but both the Navy and Texas A&M got busy, and using different approaches figured out how to clean the phages to the point they could be given safely.”
More hurdles: Legal staff at Texas A&M expressed concern about future lawsuits. “I remember the lawyer saying to me, ‘Let me see if I get this straight. You want to send unapproved viruses from this lab to be injected into a person who will probably die.’ And I said, “Yeah, that’s about it,’” Young said.
“But Stephanie literally had speed dial numbers for the chancellor and all the people involved in human experimentation at UC San Diego. After she calls them, they basically called their counterparts at A&M, and suddenly they all began to work together,” Young added.
“It was like the parting of the Red Sea — all the paperwork and hesitation disappeared.”
The purified cocktail from Young’s lab was the first to arrive in San Diego. Strathdee watched as doctors injected the Texas phages into the pus-filled abscesses in Patterson’s abdomen before settling down for the agonizing wait.
“We started with the abscesses because we didn’t know what would happen, and we didn’t want to kill him,” Schooley said. “We didn’t see any negative side effects; in fact, Tom seemed to be stabilizing a bit, so we continued the therapy every two hours.”
Two days later, the Navy cocktail arrived. Those phages were injected into Patterson’s bloodstream to tackle the bacteria that had spread to the rest of his body.
“We believe Tom was the first person to receive intravenous phage therapy to treat a systemic superbug infection in the US,” Strathdee told CNN.
“And three days later, Tom lifted his head off the pillow out of a deep coma and kissed his daughter’s hand. It was just miraculous.”
Today, nearly eight years later, Patterson is happily retired, walking 3 miles a day and gardening. But the long illness took its toll: He was diagnosed with diabetes and is now insulin dependent, with mild heart damage and gastrointestinal issues that affect his diet.
“He isn’t back surfing again, because he can’t feel the bottoms of his feet, and he did get Covid-19 in April that landed him in the hospital because the bottoms of his lungs are essentially dead,” Strathdee said.
“As soon as the infection hit his lungs he couldn’t breathe and I had to rush him to the hospital, so that was scary,” she said. “He remains high risk for Covid but we’re not letting that hold us hostage at home. He says, ‘I want to go back to having as normal life as fast as possible.’”
To prove it, the couple are again traveling the world — they recently returned from a 12-day trip to Argentina.
“We traveled with a friend who is an infectious disease doctor, which gave me peace of mind to know that if anything went sideways, we’d have an expert at hand,” Strathdee said.
“I guess I’m a bit of a helicopter wife in that sense. Still, we’ve traveled to Costa Rica a couple of times, we’ve been to Africa, and we’re planning to go to Chile in January.”
Patterson’s case was published in the journal Antimicrobial Agents and Chemotherapy in 2017, jump-starting new scientific interest in phage therapy.
“There’s been an explosion of clinical trials that are going on now in phage (science) around the world and there’s phage programs in Canada, the UK, Australia, Belgium, Sweden, Switzerland, India and China has a new one, so it’s really catching on,” Strathdee told CNN.
Some of the work is focused on the interplay between phages and antibiotics — as bacteria battle phages they often shed their outer shell to keep the enemy from docking and gaining access for the kill. When that happens, the bacteria may be suddenly vulnerable to antibiotics again.
“We don’t think phages are ever going to entirely replace antibiotics, but they will be a good adjunct to antibiotics. And in fact, they can even make antibiotics work better,” Strathdee said.
In San Diego, Strathdee and Schooley opened the Center for Innovative Phage Applications and Therapeutics, or IPATH, in 2018, where they treat or counsel patients suffering from multidrug-resistant infections. The center’s success rate is high, with 82% of patients undergoing phage therapy experiencing a clinically successful outcome, according to its website.
Schooley is running a clinical trial using phages to treat patients with cystic fibrosis who constantly battle Pseudomonas aeruginosa, a drug-resistant bacteria that was also responsible for the recent illness and deaths connected to contaminated eye drops manufactured in India.
And a memoir the couple published in 2019 — “The Perfect Predator: A Scientist’s Race to Save Her Husband From a Deadly Superbug” — is also spreading the word about these “perfect predators” to what may soon be the next generation of phage hunters.
How naturally occurring viruses could help treat superbug infections
“I am getting increasingly contacted by students, some as young as 12,” Strathdee said. “There’s a girl in San Francisco who begged her mother to read this book and now she’s doing a science project on phage-antibiotic synergy, and she’s in eighth grade. That thrills me.”
Strathdee is quick to acknowledge the many people who helped save her husband’s life. But those who were along for the ride told CNN that she and Patterson made the difference.
“I think it was a historical accident that could have only happened to Steffanie and Tom,” Young said. “They were at UC San Diego, which is one of the premier universities in the country. They worked with a brilliant infectious disease doctor who said, ‘Yes,’ to phage therapy when most physicians would’ve said, ‘Hell, no, I won’t do that.’
“And then there is Steffanie’s passion and energy — it’s hard to explain until she’s focused it on you. It was like a spiderweb; she was in the middle and pulled on strings,” Young added. “It was just meant to be because of her, I think.”